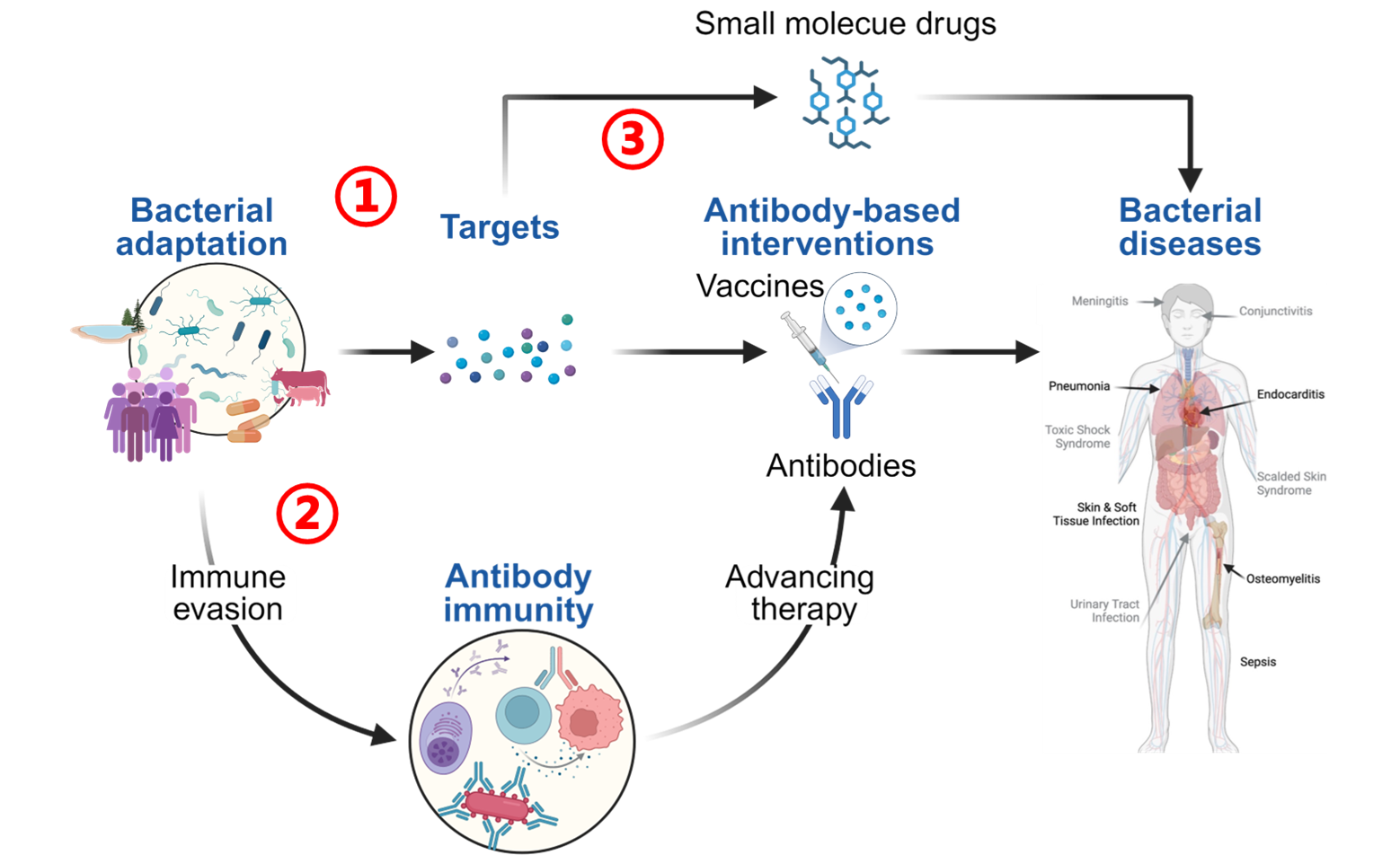
General Research Direction
Humans live in symbiosis with a diverse range of microbes that contribute to host health, yet some are capable of causing infectious diseases. Despite considerable medical progress over the past two decades, infectious diseases remain a leading cause of death globally, primarily driven by the rise of antimicrobial resistance (AMR). Importantly, many clinically significant pathogens persistently colonize the human body or environmental reservoirs, constituting a major source of subsequent infection. While antibody-based interventions, such as vaccines and monoclonal antibody therapies, hold great promise for infection control, most high-priority pathogens still lack effective options, and numerous clinical trials have faltered due to insufficient or ineffective antibody responses. Given that antibody activity during infection is shaped by pathogen, host, and structural factors, my research seeks to clarify the dynamic interplay among microbial infection, antibody function, and host immunity to inform the design of more effective antibody-based strategies. Furthermore, I aim to develop immunotherapies that not only elicit protective immunity but also restore microbial susceptibility to antibiotics. A key area of interest is investigating how prolonged use of non-antimicrobial drugs, commonly prescribed for chronic conditions, affects the host microbiota and how these changes may inadvertently promote pathogenicity and AMR. We are also pursuing the development of novel antimicrobial agents and studying bacterial adaptive mechanisms that underpin resistance evolution.
I) Main Research Area 1: Staphylococcus aureus infections cause over one million global deaths annually, making it the deadliest bacterial pathogen. The prevalence of methicillin-resistant S. aureus (MRSA), recognized as one of the most challenging multidrug-resistant organisms in healthcare settings, highlights the urgent need for novel strategies beyond traditional antibiotics. Antibody-based interventions represent a promising alternative. Nevertheless, the development of effective antibody therapies has faced significant setbacks: of the 11 antibody candidates that have entered clinical trials, 7 have already failed by 2024. The remaining four target antigens exhibit substantial overlap with those of unsuccessful candidates, raising considerable concern about their clinical potential. These persistent failures reinforce my belief that selecting effective antigen/epitope targets remains the major bottleneck, and overcoming this challenge is central to my approach in developing successful antibody interventions against S. aureus.
II) Main Research Area 2: Antibodies are critical for controlling bacterial infections. Given the growing challenge of antibiotic resistance, developing effective antibody-based interventions has become increasingly urgent. However, very few such interventions are currently approved for bacterial infections, and many candidates have encountered failures in clinical trials. A key underlying cause is our limited understanding of the mechanisms that impair antibody efficacy during infection. My research aims to address this gap by systematically investigating the factors that suppress antibody functionality in the context of bacterial infection, including pathogen-specific evasion strategies and host-mediated immunosuppressive mechanisms. The goal is to identify molecular and cellular determinants of antibody failure and leverage this knowledge to design novel antibody-based therapies that overcome these limitations and enhance immune protection.
III) Main Research Area 3: This research area begins by understanding bacterial adaptive mechanisms that underpin resistance evolution, which is critical to addressing the challenge of AMR. Building on this foundation, the project aims to develop novel therapeutic strategies, including new antimicrobial agents and potentiators that enhance the efficacy of established antibiotics such as vancomycin, daptomycin, and β-lactams. While long-term goals include designing original antibacterial compounds, the current emphasis is on rescuing and repurposing existing antibiotics through synergistic combinations. The work integrates high-throughput screening of chemical libraries, detailed mechanistic studies, and validation in both in vitro and in vivo preclinical models. Success in this endeavor could significantly extend the clinical utility of current antibiotics, mitigate the spread of multidrug-resistant infections, and contribute sustainable strategies to the global antibiotic pipeline.